Document Type : Original article
Subjects
Abstract
Background: To assess prevalence of prostate cancer and clinically significant prostate cancer in patients with PI-RADS 3 on bpMRI and factors associated with them.
Methods: Patients suspicious for prostate cancer following serum Prostate Specific Antigen (PSA) screening, who had PI-RADS 3 on bpMRI, were included. All patients underwent systemic plus MRI targeted biopsy of prostate. Gleason score 3+3 was considered positive for prostate cancer but clinically non-significant one. Higher scores were pertained to as clinically significant prostate cancer. The relationship between patient age, PSA level, PSA density, number of core biopsies, and number of PI-RADS 3 lesions on bpMRI with presence of prostate cancer per se and presence of clinically significant prostate cancer in our patients is assessed.
Results: 244 patients were enrolled. 101 patients had prostate cancer (41.4%). Out of these 101 patients, 34 (13.9% of total) had clinically significant prostate cancer. Among different factors, only PSA density was associated with both prostate cancer (OR=1.05, p=0.001) and clinically significant prostate cancer (OR=1.03, p=0.001). According to receiver operating characteristic curve analysis, best cut off value of PSA density which has highest association with clinically significant prostate cancer in PI-RADS 3 patients would be 0.36; with a sensitivity of 0.38 and specificity of 0.93.
Conclusion: Considering PSA density with threshold of 0.36 for performing biopsy in patients with PI-RADS 3 on bpMRI might lower the rate of unnecessary biopsies.
Keywords: Biparametric MRI, Clinically significant prostate cancer, PI-RADS, PSA, PSA density
Introduction
Prostate cancer is the most common solid organ malignancy among men worldwide (1,2). The current gold standard test for prostate cancer diagnosis is 12-core TRUS-guided biopsy for men with elevated serum Prostate Specific Antigen (PSA) levels or Digital Rectal Examination (DRE) abnormalities (3). Recently, Magnetic Resonance Imaging (MRI) is used to increase diagnostic accuracy. While multiparametric MRI (mpMRI) has gained acceptance as main technique of MRI for prostate cancer by using PI-RADS (Prostate Imaging Reporting and Data System) scoring, new studies show that biparametric MRI (bpMRI) has “similar PI-RADS scoring and thus diagnostic performance” (4). Prostate lesions with PI-RADS 1 or 2 are considered low likelihood of cancer and PI-RADS 4 or 5 is highly suspicious for malignancy and must undergo biopsy (5), but there is still controversy regarding actions that should be taken following PI-RADS 3, as these lesions might turn out to be either benign, clinically non-significant prostate cancer, or clinically significant prostate cancer (defined as Gleason Score >=3+4). As PI-RADS 3 is reported in 15 to 32% of MRIs of prostate (5,6), it is believed that more attention is required regarding interpreting this finding in order to postpone unnecessary biopsies.
Therefore, in this study, patients with PI-RADS 3 lesion were examined on bpMRI and checked frequency of the 3 possible pathologies that might be observed in them. Also, factors that had association with the pathology results in these patients were assessed.
Materials and Methods
Patients suspicious for prostate cancer following serum PSA screening, who had PI-RADS 3 on bpMRI, were included in the study. A serum PSA level of over 4.0 ng/ml for men older than 65 years and 2.5 ng/ml for younger men was considered to be suspicious for prostate cancer. All the patients underwent systemic plus MRI targeted biopsy of prostate. Gleason score 3+3 was considered positive for prostate cancer but clinically non-significant one. Higher scores –even in one core- were pertained to as clinically significant Prostate Cancer (csPCa), while lower scores were considered negative for csPCa.
The relationship between patient age, PSA level, PSA density (serum PSA level divided by prostate volume), number of core biopsies, and number of PI-RADS 3 lesions on bpMRI with presence of prostate cancer per se and presence of csPCa in the patients was assessed.
All the procedures were in accordance with the ethical standards of the institutional and national research committee (s) and with the Declaration of Helsinki (as revised in 2013). All the patients signed the informed consent for scientific research.
Mean±standard deviation and frequency (percentage) were used to describe the quantitative and categorical variables, respectively. Univariate and multivariate logistic regression analysis were utilized to determine the factors associated with presence of prostate cancer and csPCa. A Receiver Operating Characteristic (ROC) curve was used to define cut-off value of Prostate Specific Antigen Density (PSAD) for presence of csPCa. Data analyses were performed using STATA17 software (College Station, Texas 77845 USA). p-values less than 0.05 were considered to be statistically significant.
Results
244 patients were enrolled in the study. The mean age of the participants was 63 years. The mean amount of PSA among the patients was 7.50. Mean PSAD was 0.15. Out of 244 patients, 165 (67.6%) had one lesion, 77 (31.6%) had two lesions, and 2 patients (0.8%) had three lesions compatible with PI-RADS 3 description.
After assessing the biopsy specimens, 143 patients showed no evidence of cancerous pathologic changes (58.6%), while 101 patients had prostate cancer (41.4%). Out of 101 patients, 34 (13.9% of total) had csPCa.
Factors that might have an association with the presence of prostate cancer and clinically significant prostate cancer, were assessed. Patient age, number of PIRADS 3 lesions, number of core biopsies, and prostate volume were not associated with the presence of prostate cancer, nor with presence of csPCa.
In the univariate analysis, serum PSA level and PSAD were both associated with presence of prostate cancer and csPCa in the patients. Multivariate analysis demonstrated that only PSAD was associated with both prostate cancer (OR=1.05, p=0.001) and csPCa (OR=1.03, p=0.001).
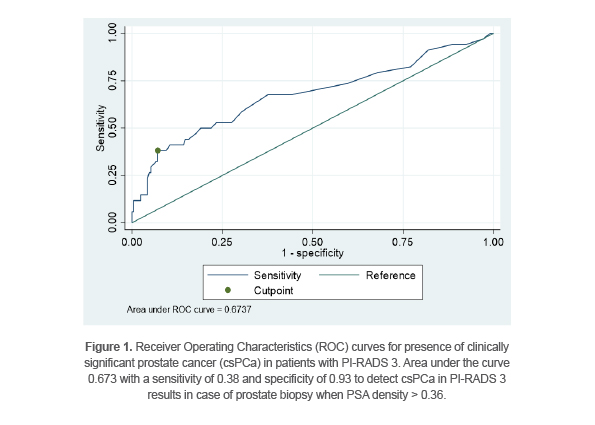
A ROC curve was used in order to define the cut-off value of PSAD for presence of csPCa, according to which, the best cut-off value of PSAD for csPCa would be 0.36 with a sensitivity of 0.38 and specificity of 0.93 (Figure 1).
Discussion
Prostate cancer is the most common solid organ malignancy among men worldwide. It is also the third leading cause of cancer-related death among men in the United States (1,2). The current gold standard test for prostate cancer diagnosis is 12-core TRUS-guided biopsy. One of the disadvantages of this diagnostic method is its susceptibility to miss prostate cancer in the peripheral zone due to the random nature of it. Also, the anterior zone is not assessed thoroughly in this procedure. The other downside of this procedure is over-diagnosis (and the resultant over-treatment), considering PSA levels as the main indication for biopsy. Thus, mpMRI is introduced to increase diagnostic accuracy for this disease. This MRI technique is already an integral part of the diagnostic work-up in brain and breast tumors, and has enhanced diagnostic process, especially by using PI-RADS scoring system (7-9). New studies show that bpMRI has same diagnostic accuracy for prostate cancer as mpMRI (4,10). Therefore, in this study, bpMRI was used in order to assess prostate lesions according to the PI-RADS scoring system.
While assessment of PI-RADS 1-2 and 4-5 are generally obvious, there is still controversy regarding interpreting PI-RADS 3 results. Venderink et al suggested that 65% of PI-RADS 3 results turned out to be benign while 35% would be cancerous, 17% of them were csPCa (11). Other studies found prostate cancer would be present in 10 to 43% of PIRADS 3 results with 4 to 21% of them being csPCa (12-15). The current study revealed that prostate cancer is observed in 41.4% of PI-RADS 3 results and 13.9% of them happens to be csPCa which is in accordance with previous studies.
Surprisingly, there are still few studies with main purpose of investigating factors associated with the presence of prostate cancer and csPCa in PI-RADS 3 results. Felker et al found that PSAD > 0.15 ng/ml2 is associated with the presence of csPCa in PI-RADS 3 (15). Bhat et al and Boesen et al also showed that high PSAD increases the chance of csPCa in PI-RADS 3 results (16,17). Hermie et al suggested a combined predictive model using prostate volume and a radiologic feature (5). Liu et al demonstrated that both prostate volume and PSAD had a significant relationship with prostate cancers independently (18).
The current study represented that only PSAD is associated with presence of prostate cancer and csPCa in PI-RADS 3 results which is in concordance with the previous ones. While above mentioned studies considered the limit of 0.15 ng/ml2 for PSAD as suggested by Epstein et al (19), a Receiver Operation Characteristic (ROC) curve analysis was performed in the present study to assess if another value for PSAD could be more suitable for finding csPCa in PI-RADS 3 patients. Considering the area under curve, it is suggested that the best cut-off point for PSAD in PI-RADS 3 patients should be 0.36 (with a sensitivity of 0.38 and specificity of 0.93 to be associated with csPCa) (Figure 1). Considering this cut-off value, it is suggested in patients with PI-RADS 3, biopsy should be offered if PSAD is higher than 0.36; in those with PSAD values below 0.36, follow up might be more suitable option.
This research was subjected to multiple limitations, since it was performed as a retrospective study in a single center. As we know, rate of csPCa in PI-RADS 3 patients is less than 50%, thus multi-center registry-based studies may include more cases of csPCa which makes the results more replicable. Considering the location of pathologic lesions in further studies would also be preferable as different PI-RADS 3 lesions may have different characteristics based on their locations on MRI.
Conclusion
Considering PSAD with threshold of 0.36 for performing biopsy in patients with PI-RADS 3 on bpMRI might lower the rate of unnecessary biopsies and its possible complications.
Funding
None.
Acknowledgement
The ethical code number of the Ethics committee for this study is IR.IUMS.REC.1403.231.
Disclosure
None
Conflict of Interest
There was no conflict of interest in this manuscript.